In order to increase the efficiency of NMR-based protein-ligand binding screens, small molecules are often screened as mixtures. Generally, mixture sizes have been arbitrarily chosen to contain 5-10 compounds. An analysis of the efficiency of mixtures and a corresponding determination of the optimum mixture size (OMS) that minimizes the amount of material and instrumentation time required for an NMR screen has been lacking. A model for calculating OMS based on the application of the hypergeometric distribution function to determine the probability of a "hit" for various mixture sizes and hit rates has been determined.
Hypergeometric distribution function
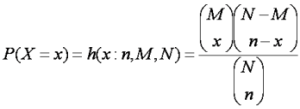
Where P is the probability of a mixture containing at least one hit, x is the number of hits present in a mixture, n is the number of compounds in the mixture, M is the number of hits present in the library, and N is the total population of the library.
Impact of Deconvolution
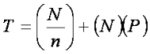
Where T is the total number NMR experiments collected to identify binders in a library.
The OMS is strongly hit-rate dependent and drops dramatically as the hit-rate approaches typical values (~5%) observed for directed or focused chemical libraries. Since the efficiency of a screen (minimal number of NMR experiments) is always greater in the absence of deconvolution, it is always better to assay a directed library using small mixture sizes (3-4) that does not require deconvolution.
References
- (47) K. A. Mercier and R. Powers* (2005) "Determining the Optimal Size of Small Molecule Mixtures for High Throughput NMR Screening", J. Biomol. NMR, 31(3):243-258. PMID15803397.
Picture Gallery
- From: Journal of Biomolecular NMR (2005)