The Function of a Protein is Intrinsically Linked to the Identity of its Active-Site and Ligand
FAST-NMR combines structural biology and NMR ligand affinity screens with bioinformatics to assign a function to a hypothetical protein or a protein of unknown function. This is based on basic tenets of biochemisttry where proteins with similar functions will have interactions are determined through a tierd NMR screen using a library composed of compounds with known biological activity. A rapid protein-ligand co-structure is determined by combining the experimental identification of the ligand-binding site from NMR chemical shift perturbations with the protein-ligand docking program AutoDock. Our CPASS (Comparison of Protein Active Site Structures) software and database is then used to compare this active site with proteins of known function.
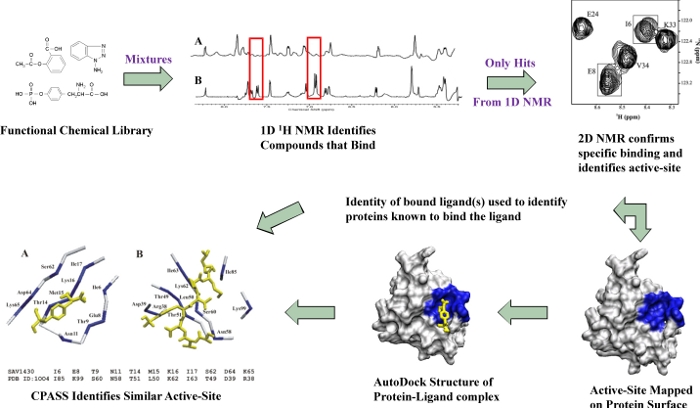
Our functional chemical library is first screened as mixtures using a 1D NMR line-broadening screen to rapidly identify binders. The relative change in peak intensity or line-width is used to measure a semi-quantitative dissociation constant (KD) that allows the ligands to be ranked. Only the ligands that exhibit a positive binding interaction in the 1D NMR line-broadening screen are further screened as singletons by measuring chemical shift changes in 2D 1H-15N HSQC spectra.
The observation of a consensus clustering of residues that experienced a chemical shift perturbation (CSP) on the molecular surface of the protein confirms a specific, stoichiometric binding interation for the ligand and the ligand-defined binding site or the proteins active-site/functional-site.
The CSPs are used in combination with AutoDock to rapidly determine (~30-45 min.) a protein-ligand co-strucure. The co-structure is then used as input for CPASS to identify potential functional homologs based on the sequence and structure similarity of ligand-defined binding sites (higher information content than global sequence or structure similarity searches).
Approximately 50% of the genome of multiple organisms contain proteins of unknown function, where 30-50% of these proteins are amenable to NMR analysis.
In cases where a structure and/or NMR assignments of a hypothetical protein is unavailable, FAST-NMR can provide functional information by the similarity in the ligand-binding profile. Comparable to sequence homology, the relative KD measured for each ligand against a protein of unknown function can be compared against this same list of binding affinities for proteins of known function. A similarity score can be obtained based on the relative differences in the individual KD values. A function can then be implied based on a high similarity in the ligand binding profiles.
Also, the output of FAST-NMR is a protein-ligand co-structure, which may provide the starting point for a structure-based drug design effort. This is especially true since the compounds in the functional chemical library have "drug-like" characteristics.
References
- (58) K. A. Mercier, M. Baran, V. Ramanathan, P. Revesz, R. Xiao, G. T. Montelione and R. Powers* (2006) "FAST-NMR - Functional Annotation Screening Technology Using NMR", J. Amer. Chem. Soc., 128(47):15292-15299. PMC2529462.
- (62) R. Powers*, J. Copeland and K. Mercier (2008) "The Application of FAST-NMR for the Identification of Novel Drug Discovery Targets", Drug Discov. Today, 13(3-4):172-179. PMC18275915.
- (61) R. Powers* (2007) "Functional Genomics and NMR Spectroscopy", Comb. Chem. High Throughput Screening, 10(8):676-697. PMID18045080.
Picture Gallery
- From: JACS (2006)
- Process Flow Chart of FAST-NMR
- Functional assignments for hypothetical protein SAV1430 from S. aureus
- Sequence alignment of SAV1430 with NifU
- Hypothetical Proteins from Structural Genomics
- From: Drug Discovery Today (2008)